Set the selected DNA or amino acid sequence to one of ten colors.Ĭolor two DNA strands or protein sequences. When copying and pasting the sequence, the function is automatically transferred.Įdit the ends of linear DNA sequences to add or remove overhangs or phosphates. The ancestral protein or DNA sequence can be reproduced as a separate file. View the automatically generated protein sequence graph history. The checkbox toggles between compact or fully expanded display. Sort the annotated feature list by name, location, size, color or type. The same analysis can be performed on selected parts of the protein. View protein molecular weight, extinction coefficient, isoelectric point and amino acid composition. Use 1- or 3-letter amino acid codes with related characteristics to view protein sequences. View multiple views of protein sequences.Ĭustomize the display of areas, sites, keys and sequence colors. Use the multi-function controls to adjust the zoom factor and display area. Use the proprietary MICA algorithm to find the sequence in the chromosome immediately. Use SnapGene's efficient data processing function to scan large DNA sequences with thousands of annotation functions. Ancestor sequence can be restored as a separate file View the automatically generated graphical history of DNA constructs. Sort the hybrid primer list by name, length, color, binding site, directionality or melting temperature. Sort the annotated feature list by name, location, size, color, directionality or type. Cut sites can be displayed as numbers or rows, sorted by name or frequency. Single chain mode displays a compact overview with color features.Ĭhoose from a series of enzyme groups. Use double-stranded mode to view enzyme sites, with features of translation, primers, and DNA color. The map can be in circular or linear format. This will determine whether or not it will be read correctly.Customize the display of enzyme sites, features, primers, ORF, DNA color, etc. In addition, the combination of restriction enzymes you use determines how a genetic sequence will fit in a construct. 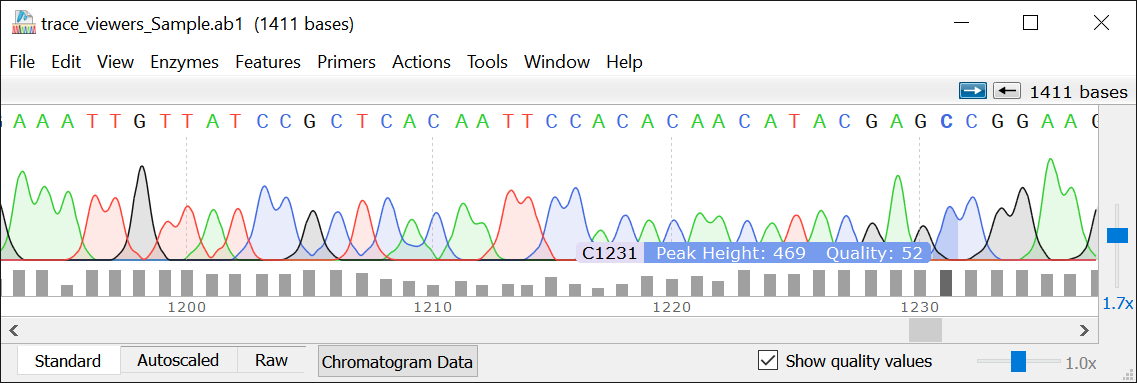
This is of particular importance because you don’t want to use an enzyme that can cut somewhere in the middle of a genetic sequence. 
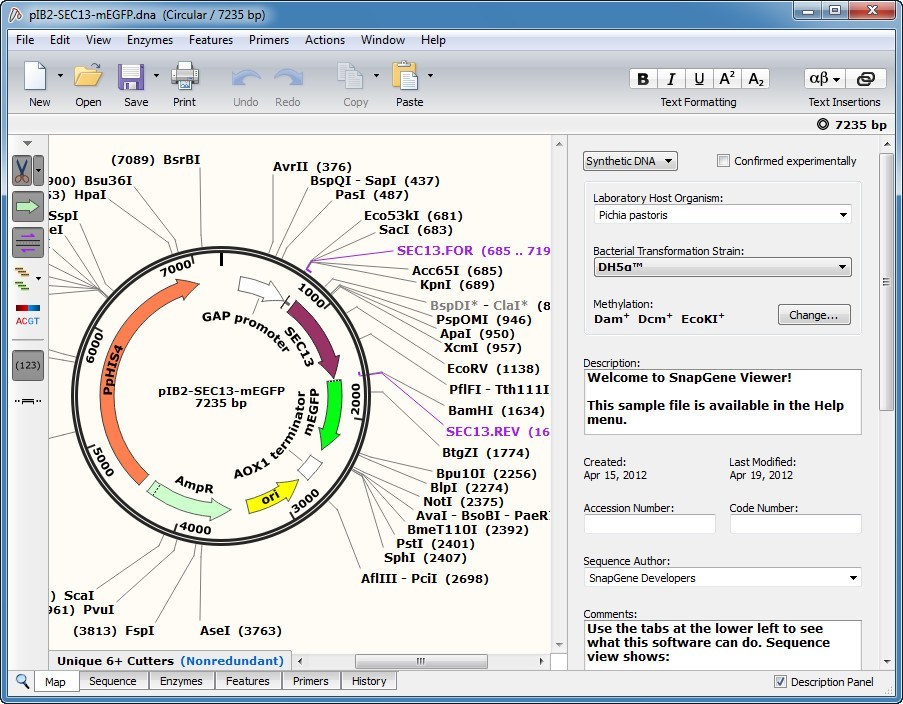
There are a number of things to consider when applying restriction cloning, such as which restriction enzymes to use. SnapGene allows you to simulate this process either by using the protocols it has available or creating your own. This process is performed in the lab following a given protocol. A number of cut sequences can be joined together with a process known as ligation. Restriction cloning is a method of editing genetic sequences by cutting them with restriction enzyme at suitable restriction sites. 
The Sequence view also shows the features seen in the map view, and adds an extra feature, a red mark that indicates the stop codons of your sequence. The Sequence view shows the characters in the sequence is convenient when editing sequences because it allows you to pinpoint the critical locations in the sequence. This information is important for estimating the cost of synthesizing your sequence and is useful in a number of calculations. This includes the number of base pairs, the number of amino acids and the molecular weight of the sequence in Daltons. The Features view, which is accessed by clicking on Views and selecting Features, shows some information about the sequences you have. These views allow you to see features included in your file, the restriction enzymes in your sequence, the actual characters in the sequence, and your primers. Other views can be accessed by selecting one of the options in the Views menu. If the length and direction of the reading frames feature are the same as the length and direction of your gene’s feature, the gene will likely be transcribed correctly. These features allow you to see the reading frames of the sequences you have in your file, and highlights the parts of your construct that hold given sequences.Ĭlicking on show translate on the left adds some features that show the reading frames of each sequence. The default view is the map which hides the letters in the genetic sequence to create space for visual features. There are a number of ways you can view a genetic sequence in SnapGene. This is extremely useful in a field like synthetic biology where existing sequences are combined to form new ones that allow organisms to perform novel functions. SnapGene is software that allows you to visualize and edit genetic sequences. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
